'Recent data suggests that more than 41% of the cases found in India belong to A3i while the most dominant clade is A2a which accounts for more than 50% of the cases.'
'This shows A3i is the second most dominant strain.'
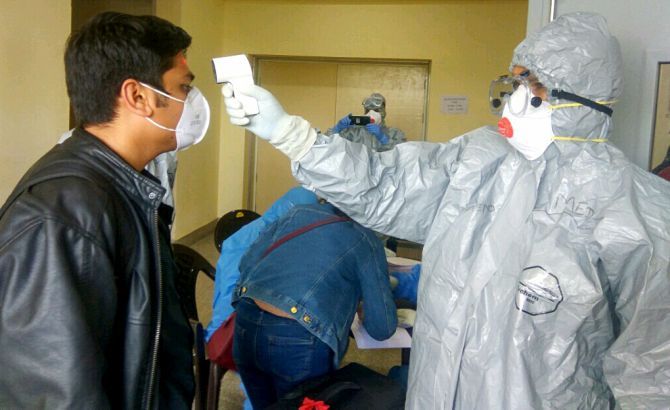

Scientists at the Centre for Cellular and Molecular Biology, Hyderabad, discovered a new strain of the novel coronavirus, Clade A3i, that has already infected around 41% of the cases in India.
The CCMB scientists have found that A3i came not from China or Europe, but from South-East Asia.
"In India, A3i is found mostly in southern India, and a few in Delhi. You don't see this strain much in, say, Gujarat or Maharashtra," Dr Rakesh K Mishra, director, CCMB, tells Shobha Warrier/Rediff.com.
The first of a two-part interview:
How significant is the discovery of the new strain of the novel coronavirus, Clade A3i?
Clade A3i is unique among the already known strains of coronavirus as A3i is defined by four mutations.
What we have found is, this has come to India, probably to the south of India, in the middle of February from South-East Asia. From there it has spread to the rest of India.
Recent data suggests that more than 41% of the cases found in India belong to A3i while the most dominant clade is A2a which accounts for more than 50% of the cases.
This shows A3i is the second most dominant strain.
It was earlier reported that there were 10 clades of coronavirus and the dominant one was A2a. Does that mean A3i is different from all the 10 clades?
Yes. This is a new strain that we have found. And this strain is seen predominantly in the South East Asian region like India, Singapore, Vietnam, Brunei, Philippines, etc.
This strain is not seen in Europe or China. It looks like it reached India in mid to late February from this region.
In India also, A3i is found mostly in the southern parts of India, and a few in Delhi.
You don't see this strain much in, say, Gujarat or Maharashtra.
It appears that A2a, though it came later, spreads faster. It also could be because of the movement of people.
There are two major factors that cause the fast spread of the virus; one is the virulence of the virus, and the other is the movement of people as it spreads from person to person.
Is mutating faster something special to coronavirus? Or, do all viruses mutate like this?
Most viruses mutate pretty fast.
In fact, SARS-CoV-2 mutates slower compared to other viruses, but it is still a pretty high number -- roughly once in every two weeks.
However, what we have noticed in our study is that the mutation rate of A3i is slower than the other strains like A2a.
Does that mean as the virus moved from Wuhan to South-East Asia, it mutated many times and became this new strain?
 Exactly. Coronavirus mutates once every two weeks, and you don't find the B virus that originated in Wuhan here.
Exactly. Coronavirus mutates once every two weeks, and you don't find the B virus that originated in Wuhan here.
In fact, in these parts you don't find B at all; you only see A2a, A3i, etc.
The advantage of the knowledge about this distinct clade is that we can follow it and find out the different locations where it is found.
We can then understand how it spreads; that is, from which city to which city. We can very precisely map the path of the virus.
So, finding of this clade gives us the correct insight into the spread of the virus.
Does a virus become slower in spreading as it mutates several times?
Parasites normally change as they evolve.
Typically, as a virus spreads and mutates, it becomes less efficient with milder symptoms.
If the virus is extremely lethal, it will not spread beyond a point as the infected people die too quickly.
In this case, what we have found in the mutations is that three are non-synonymous mutations.
What it means is, it changes the protein sequence to something different from the original.
Out of these three, one mutation is interesting because it is located in the region that is important for the virus to copy itself.
Computational analyses indicate that this mutation may possibly reduce the efficiency with which the virus can copy itself, though we have to do specific experiments to validate this.
When we say the virus is less efficient, it means it can be less virulent, low in showing symptoms, asymptomatic, etc.
If it turns out to be true, it is good news for us.
The other day, ICMR said that 80% of the cases in India are asymptomatic now. Is it because of non-synonymous mutation?
It is impossible to say that with certainty without enough clinical data. There may be several other factors contributing to that, and this mutation could be one of them, if at all.
In other countries also, there are a large number of asymptomatic patients. Yes, it looks like they are more in India.
Maybe the virus is less virulent, maybe people are more efficient or maybe a mild version of a closely related virus was here and people may have developed immunity.
We have to do a large study on that to get a clearer picture.
Yes, it shows this virus is somewhat sluggish compared to other countries. We are now collecting samples from different places at different points of time.
For example, in Telangana what was most common earlier was A3i, but now we see a lot more A2a cases coming up.
It may be because A3i was slow in spreading, and slowly A2a came here and started spreading faster.
Which is more dangerous, A2a or A3i as far as mortality is concerned?
There is no data available to show that mortality depends on clade.
We are yet to get that data because sequencing is done by different groups and mortality takes place in hospitals.
In fact, we are collecting data from various hospitals. In a couple of weeks, we will have some idea if there is any connection between mortality and clade type.
However, data from other countries on the existing 10 clades so far does not show correlation between clade and mortality.
Now that we know there are two major clades, there are many interesting questions to answer.
If we are able to find out whether one clade is more symptomatic and the other one is less, then we can find out the biology of the virus; the genome of the virus.